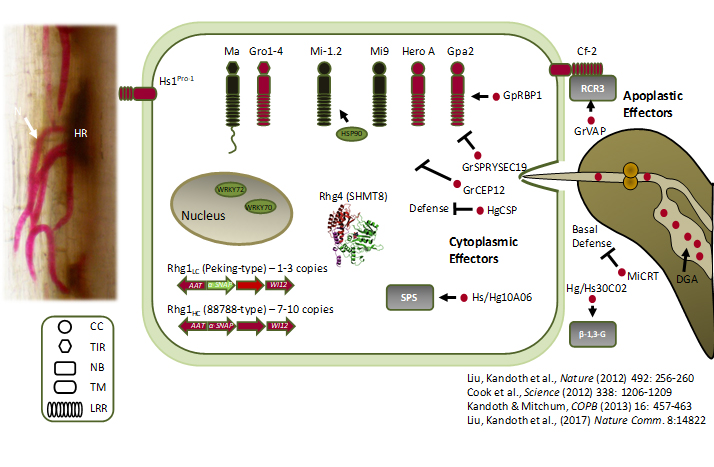
Our current understanding of how plants resist infection by plant-parasitic nematodes is limited. Only a handful of plant resistance genes to nematodes have been identified and even less is known about the actual mechanism of resistance to these important pests. In soybean, nematodes penetrate the roots of susceptible and resistant varieties equally well; however, feeding cells degenerate in resistant plants and the immobilized nematodes starve to death. Despite the plant’s best efforts, nematodes are able to overcome host resistance and reproduce on resistant varieties. The identity of SCN and RKN virulence genes that confer virulence on soybean remains unknown.
A major focus of research in my lab has been directed at understanding the molecular basis of plant resistance to SCN and RKN. We have employed laser microdissection, a technique that allows for the isolation of a single cell type from a complex mixture of cells or tissues to isolate feeding cells from resistant and susceptible plants differing by a single resistance gene. This led to the identification of genes potentially involved in mediating the plant’s resistance mechanism to the nematode. Moreover, efforts by my lab led to the discovery of a soybean gene with a major role in resistance to SCN. The resistance gene at the Rhg4 (for resistance to Heterodera glycines) locus on chromosome 8, codes for the enzyme soybean serine hydroxymethyltransferase (GmSHMT8), a key cytosolic enzyme in cellular 1-carbon folate metabolism. This discovery represented the first report of SHMT in plant resistance and has uncovered a potentially novel mechanism of resistance to nematodes. Additionally, efforts by my lab have led to the discovery of additional GmSNAP genes involved in resistance to SCN, including the GmSNAP02 gene, that functions in resistance through a loss-of-function mechanism. Recently, the RKN resistance locus Rmi1 was fine-mapped to two genes on chromosome 10, a glycosyl hydrolase family 9 β-1,4-endoglucanase gene and a type I pectin methylesterase gene. Understanding the resistance mechanisms mediated by these genes to confer resistance to SCN and RKN is currently an active area of research in my lab. Current work includes RNAseq, metabolite profiling, CRISPR gene editing, protein-protein interactions, and collaborative efforts on the structural and biochemical properties of the resistance proteins.
Furthermore, to explore the molecular basis of nematode virulence, we have been selecting nematode populations adapted to resistant varieties. These populations are being exploited for genome and transcriptome comparisons to identify genes underlying nematode virulence.
Related Publications:
Howland A, Monnig N, Mathesius J, Nathan M, Mitchum MG. Survey of Heterodera glycines population levels and virulence phenotypes during 2015-2016 in Missouri. Plant Disease 2018;102:2407-2410. https://doi.org/10.1094/PDIS-04-18-0650-SR
Shannon G, Nguyen HT, Crisel M, Smothers S, Clubb M, Vieira CC, Ali ML, Selves S, Mitchum MG, Scaboo A, Li Z, Bond J, Meinhardt C, Robbins RT, Chen P. Registration of ‘S11-20124C’ soybean with high yield potential, multiple nematode resistance, and salt tolerance. Journal of Plant Registrations 2018; 13(2):154-160. https://doi.org/10.3198/jpr2018.06.0041crc
Shannon G, Crisel M, Smothers S, Ali ML, Clubb M, Selves S, Mitchum MG, Scaboo A, Li Z, Bond J, Meinhardt C, and Chen P. Registration of ‘MO 5301D CONV’ Soybean. Journal of Plant Registrations 2018; 13(2):148-153. https://doi.org/10.3198/jpr2018.02.0006crc
Kandoth PK, Liu S, Prenger E, Ludwig A, Lakhssassi N, Heinz R, Zhou Z, Howland A, Gunther J, Warren S, Dhroso A, LaFayette P, Tucker D, Johnson S, Anderson J, Alaswad A, Cianzio SR, Parrott WA, Korkin D, Meksem K, Mitchum MG. Systematic mutagenesis of serine hydroxymethyltransferase reveals an essential role in nematode resistance. Plant Physiology 2017; 175:1370-1380. https://doi.org/10.1104/pp.17.00553
Liu S, Kandoth P, Lakhssassi N, Kang J, Colantonio VN, Heinz R, Yeckel G, Zhou Z, Bekal S, Dapprich J, Rotter B, Cianzio S, Mitchum MG, Meksem K. The soybean GmSNAP18 underlies two types of resistance to soybean cyst nematode. Nature Communications 2017: 8:14822 https://doi.org/10.1038/ncomms14822
Gardner MN, Heinz R, Wang J, and Mitchum MG. Genetics and adaptation of soybean cyst nematode to broad spectrum soybean resistance. G3: Genes, Genomes, Genetics 2017; 7(3) 835-841; https://doi.org/10.1534/g3.116.035964
Mitchum MG. Soybean resistance to the soybean cyst nematode Heterodera glycines: an update. Phytopathology 2016;106(12):1444-1450. https://doi.org/10.1094/PHYTO-06-16-0227-RVW
Kandoth PK and Mitchum MG. War of the worms: how plants fight underground attacks. Current Opinion in Plant Biology 2013;16: 457-463. https://doi.org/10.1016/j.pbi.2013.07.001
Kandoth P, Heinz R, Yeckel G, Gross NW, Juvale PS, Hill J, Whitham SA, Baum TJ, Mitchum MG. A virus-induced gene silencing method to study soybean cyst nematode parasitism in Glycine max. BioMed Central Research Notes 2013;6:255. https://dx.doi.org/10.1186%2F1756-0500-6-255
*Liu S, *Kandoth PK, Warren SD, Yeckel G, Heinz R, Alden J, Yang C, Jamai A, El-Mellouki T, Juvale PS, Hill J, Baum TJ, Cianzio S, Whitham SA, Korkin D, †Mitchum MG, and †Meksem K. A soybean cyst nematode resistance gene points to a new mechanism of plant resistance to pathogens. Nature 2012;492:256-260. *co-first authors; †co-senior authors https://doi.org/10.1038/nature11651
Juvale PS, Hewezi T, Zhang C, Kandoth PK, Mitchum MG, Hill JH, Whitham SA, and Baum TJ. Temporal and spatial Bean pod mottle virus-induced gene silencing in soybean. Molecular Plant Pathology 2012;13(9):1140-1148. https://doi.org/10.1111/j.1364-3703.2012.00808.x
Kandoth PK, Ithal N, Recknor J, Maier T, Nettleton D, Baum TJ, and Mitchum MG. The soybean Rhg1 locus for resistance to the soybean cyst nematode Heterodera glycines regulates expression of a large number of stress- and defense-related genes in degenerating feeding cells. Plant Physiology 2011;155:1960-1975. *co-first authors https://doi.org/10.1104/pp.110.167536
Escobar C, Horowitz SB, and Mitchum MG. Transcriptomic and proteomic analysis of the plant response to nematode infection. Chapter 9 IN: J Jones, G Gheysen, and C Fenoll (Eds.) Genomics and Molecular Genetics of Plant-Nematode Interactions (pp. 157-173). New York: Springer, 2011. https://link.springer.com/chapter/10.1007/978-94-007-0434-3_9
Ithal N and Mitchum MG. Laser capture microdissection of nematode feeding cells, Chapter 18. IN: J McDowell (Ed.), Plant Immunity: Methods and Protocols, Methods in Molecular Biology, vol. 712 (pp. 227-240). New York: Springer. 2011. https://doi.org/10.1007/978-1-61737-998-7_18
*Liu X, *Liu S, Jamai A, Bendahmane A, Lightfoot D, Mitchum MG and Meksem K. Soybean cyst nematode resistance in soybean is independent of the Rhg4 locus LRR-RLK Functional and Integrative Genomics 2011;11(4):539-549. *co-first authors https://doi.org/10.1007/s10142-011-0225-4
*Brown S, *Yeckel G, *Heinz R, Clark K, Sleper D, and Mitchum MG. A high-throughput automated technique for counting females of Heterodera glycines using a fluorescence-based imaging system. Journal of Nematology 2010;42(3):201-206. *co-first authors https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3380484/
Mitchum MG, Wrather JA, Heinz RD, Shannon JG and Danekas G. Variability in distribution and virulence phenotypes of Heterodera glycines in Missouri during 2005. Plant Disease 2007;91(11):1473-1476. https://doi.org/10.1094/PDIS-91-11-1473
Mitchum MG and Baum TJ. Genomics of the soybean cyst nematode-soybean interaction. IN: G. Stacey (Ed.), Genetics and Genomics of Soybean (pp. 321-341). New York: Springer, 2008. https://link.springer.com/content/pdf/10.1007/978-0-387-72299-3_17.pdf
El-Mellouki T, Liu S, Liu X, Jamai A, Mitchum MG and Meksem K. TILLING: A reverse genetics and functional genomics tool in soybean. IN: G Kahl and K Meksem (Ed.), The Handbook of Plant Functional Genomics (pp. 251-265). Weinheim: Wiley-Blackwell, 2008 https://doi.org/10.1002/9783527622542.ch12
Ithal N, Recknor J, Nettleton D, Maier T, Baum TJ and Mitchum MG. Developmental transcript profiling of cyst nematode feeding cells in soybean roots. Molecular Plant-Microbe Interactions 2007;20(5):510-525. https://doi.org/10.1094/MPMI-20-5-0510
Ithal N, Recknor J, Nettleton D, Hearne L, Maier T, Baum TJ and Mitchum MG. Parallel genome-wide expression profiling of host and pathogen during soybean cyst nematode infection of soybean. Molecular Plant-Microbe Interactions 2007;20(3):293-305. https://doi.org/10.1094/MPMI-20-3-0293
